Aggregate classifier statistics calculator
- class simba.data_processors.agg_clf_calculator.AggregateClfCalculator(config_path: Union[str, PathLike], classifiers: List[str], data_dir: Optional[Union[str, PathLike]] = None, detailed_bout_data: bool = False, transpose: bool = True, first_occurrence: bool = True, event_count: bool = True, total_event_duration: bool = True, pct_of_session: bool = True, mean_event_duration: bool = True, median_event_duration: bool = True, mean_interval_duration: bool = True, median_interval_duration: bool = True, frame_count: bool = False, video_length: bool = False, save_dir: Optional[Union[str, PathLike]] = None)[source]
Compute aggregate descriptive statistics from classification data.
This class analyzes machine learning classifier results to calculate various descriptive statistics such as bout counts, durations, intervals, and first occurrences for each classifier in each video. Results can be saved in detailed or summary formats.
See also
For multi core class, see
simba.data_processors.agg_clf_calculator.AggregateClfCalculator()- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
classifiers (List[str]) – List of classifier names to calculate aggregate statistics for. Must be valid classifier names from the project.
data_dir (Optional[Union[str, os.PathLike]]) – Directory containing the machine results CSV files. If None, uses
project_folder/csv/machine_results.detailed_bout_data (bool) – If True, saves detailed bout data (start frame, end frame, bout time, etc.) for each bout in each video. Default: False.
transpose (bool) – If True, creates output with one video per row. If False, one measurement per row. Default: False.
first_occurrence (bool) – If True, calculates first occurrence time for each classifier. Default: True.
event_count (bool) – If True, calculates total number of bouts for each classifier. Default: True.
total_event_duration (bool) – If True, calculates total duration of all bouts for each classifier. Default: True.
mean_event_duration (bool) – If True, calculates mean duration of bouts for each classifier. Default: True.
median_event_duration (bool) – If True, calculates median duration of bouts for each classifier. Default: True.
mean_interval_duration (bool) – If True, calculates mean interval between bouts for each classifier. Default: True.
median_interval_duration (bool) – If True, calculates median interval between bouts for each classifier. Default: True.
frame_count (bool) – If True, includes total frame count in the output. Default: False.
video_length (bool) – If True, includes video length in seconds in the output. Default: False.
save_dir (Optional[Union[str, os.PathLike]]) – Optional directory in which to save the output CSVs (
data_summary_*.csvand, if requested,detailed_bout_data_summary_*.csv). If None, defaults to the project’slogs_path. Default: None.
- Raises
NoChoosenMeasurementError – If no measurement types are selected (all measurement booleans are False).
- Example
>>> clf_calculator = AggregateClfCalculator( ... config_path="project_folder/project_config.ini", ... classifiers=['Attack', 'Sniffing'], ... detailed_bout_data=True, ... transpose=True ... ) >>> clf_calculator.run() >>> clf_calculator.save()
Interpolate pose-estimation data
- class simba.data_processors.interpolate.Interpolate(config_path: Union[str, PathLike], data_path: Union[str, PathLike, List[Union[str, PathLike]]], type: Optional[typing_extensions.Literal['body-parts', 'animals']] = 'body-parts', method: Optional[typing_extensions.Literal['nearest', 'linear', 'quadratic']] = 'nearest', multi_index_df_headers: Optional[bool] = False, copy_originals: Optional[bool] = False)[source]
Interpolate missing body-parts in pose-estimation data. “Missing” is defined as either (i) when a single body-parts is None, or when all body-parts belonging to an animal are identical (i.e., the same 2D coordinate or all None).

Note
Important
The interpolated data overwrites the original data on disk. If the original data is required, pass
copy_originals = Trueto save a copy of the original data.- Parameters
config_path (Union[str, os.PathLike]) – path to SimBA project config file in Configparser format.
data_path (Union[str, os.PathLike]) – Path to a directory, path to a file, or a list of file paths to files with pose-estimation data in CSV or parquet format.
type (Optional[Literal['body-parts', 'animals']]) – If ‘animals’, then interpolation is performed when all body-parts belonging to an animal are identical (i.e., the same 2D coordinate or all None). If ‘body-parts` then all body-parts that are None will be interpolated. Default: body-parts.
method (Optional[Literal['nearest', 'linear', 'quadratic']]) – If ‘animals’, then interpolation is performed when all body-parts belonging to an animal are identical (i.e., the same 2D coordinate or all None). If ‘body-parts` then all body-parts that are None will be interpolated. Default: body-parts.
multi_index_df_headers (Optional[bool]) – If truth-like, then the input data is anticipated to have multiple header columns, and output columns will have multiple header columns. Default: False.
copy_originals (Optional[bool]) – If truth-like, then the pre-interpolated, original data, will be bo stored in a subdirectory of the original data. The subdirectory is named according to the type of interpolation and datetime of the operation.
- Example
>>> interpolator = Interpolate(config_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', data_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/csv/input_csv/test', type='body-parts', multi_index_df_headers=True, copy_originals=True) >>> interpolator.run()
Advanced pose-estimation interpolation
- class simba.data_processors.advanced_interpolator.AdvancedInterpolator(data_path: Union[str, PathLike], settings: Dict[str, Any], type: Optional[typing_extensions.Literal['animal', 'body-part']] = 'body-part', verbose: Optional[bool] = True, config_path: Optional[Union[str, PathLike]] = None, save_dir: Optional[Union[str, PathLike]] = None, multi_index_data: Optional[bool] = False, save_copy: Optional[bool] = True, max_interpolation_length: Optional[int] = None)[source]
Interpolation method that allows different interpolation parameters for different animals or body-parts. For example, interpolate some body-parts of animals using linear interpolation, and other body-parts of animals using nearest interpolation.
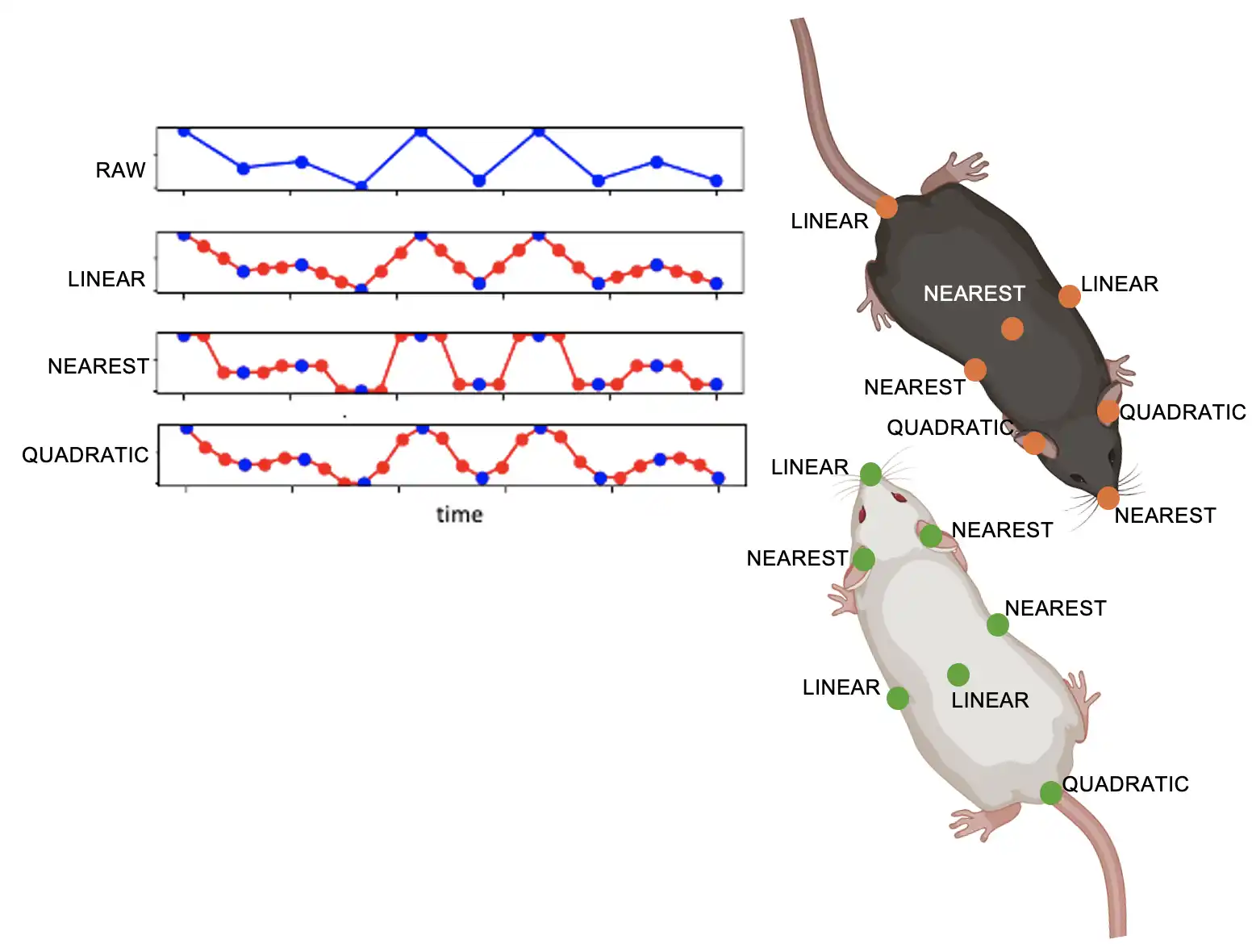
- Parameters
data_path (Union[str, os.PathLike]) – Path to folder containing pose-estimation data or a file with pose-estimation data.
config_path (Union[str, os.PathLike]) – Optional path to SimBA project config file in Configparser format.
type (Literal["animal", "body-part"]) – Type of interpolation: animal or body-part. Default: ‘body-part’.
settings (Dict) – Interpolation rules for each animal or each animal body-part. See examples.
verbose (bool) – If True, prints progress messages. Default: True.
save_dir (Union[str, os.PathLike]) – Optional directory to save results. If None, saves in input directory.
multi_index_data (bool) – If True, the incoming data has multi-index columns. Default: False.
save_copy (bool) – If True, saves original data in datetime-stamped sub-directory. Default: True.
max_interpolation_length (Optional[int]) – Maximum length of gaps to interpolate. If None, interpolates all gaps. Default: None.
- Examples
>>> # Animal-level interpolation >>> interpolator = AdvancedInterpolator( ... data_path='/path/to/project_folder/csv/input_csv', ... config_path='/path/to/project_folder/project_config.ini', ... type='animal', ... settings={'Animal_1': 'linear', 'Animal_2': 'quadratic'}, ... multi_index_data=True ... ) >>> interpolator.run() >>> >>> # Body-part level interpolation >>> interpolator = AdvancedInterpolator( ... data_path='/path/to/project_folder/csv/input_csv', ... config_path='/path/to/project_folder/project_config.ini', ... type='body-part', ... settings={ ... 'Simon': { ... 'Ear_left_1': 'linear', ... 'Ear_right_1': 'linear', ... 'Nose_1': 'quadratic', ... 'Lat_left_1': 'quadratic', ... 'Lat_right_1': 'quadratic', ... 'Center_1': 'nearest', ... 'Tail_base_1': 'nearest' ... }, ... 'JJ': { ... 'Ear_left_2': 'nearest', ... 'Ear_right_2': 'nearest', ... 'Nose_2': 'quadratic', ... 'Lat_left_2': 'quadratic', ... 'Lat_right_2': 'quadratic', ... 'Center_2': 'linear', ... 'Tail_base_2': 'linear' ... } ... }, ... multi_index_data=True ... ) >>> interpolator.run()
Smooth pose-estimation data
- class simba.data_processors.smoothing.Smoothing(config_path: Union[str, PathLike], data_path: Union[str, PathLike, List[Union[str, PathLike]]], time_window: int, method: Optional[typing_extensions.Literal['gaussian', 'savitzky-golay']] = 'Savitzky-Golay', multi_index_df_headers: Optional[bool] = False, copy_originals: Optional[bool] = False)[source]
Smooth pose-estimation data according to user-defined method.

Note
Important
The smoothened data overwrites the original data on disk. If the original data is required, pass
copy_originals = Trueto save a copy of the original data.- Parameters
config_path (Union[str, os.PathLike]) – path to SimBA project config file in Configparser format.
data_path (Union[str, os.PathLike, List[Union[str, os.PathLike]]]) – Path to directory containing pose-estimation data, to a file containing pose-estimation data, or a list of paths containing pose-estimation data.
time_window (int) – Rolling time window in millisecond to use when smoothing. Larger time-windows and greater smoothing.
method (Optional[Literal["gaussian", "savitzky-golay"]]) – Type of smoothing_method. OPTIONS:
gaussian,savitzky-golay. Default gaussian.multi_index_df_headers (bool) – If True, the incoming data is multi-index columns dataframes. Default: False.
copy_originals (bool) – If truth-like, then the pre-smoothened, original data, will be bo stored in a subdirectory of the original data. The subdirectory is named according to the type of smoothing method and datetime of the operation.
- References
- Examples
>>> smoother = Smoothing(data_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/csv/input_csv/Together_1.csv', config_path=r'/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', method='Savitzky-Golay', time_window=500, multi_index_df_headers=True, copy_originals=True) >>> smoother.run()
Advanced smooth pose-estimation data
- class simba.data_processors.advanced_smoothing.AdvancedSmoother(data_path: Union[str, PathLike], config_path: Union[str, PathLike], settings: Dict[str, Any], type: Optional[typing_extensions.Literal['animal', 'body-part']] = 'body-part', verbose: Optional[bool] = True, multi_index_data: Optional[bool] = False, overwrite: Optional[bool] = True)[source]
Smoothing method that allows different smoothing parameters for different animals or body-parts. For example, smooth some body-parts of animals using Savitzky-Golay smoothing, and other body-parts of animals using Gaussian smoothing.
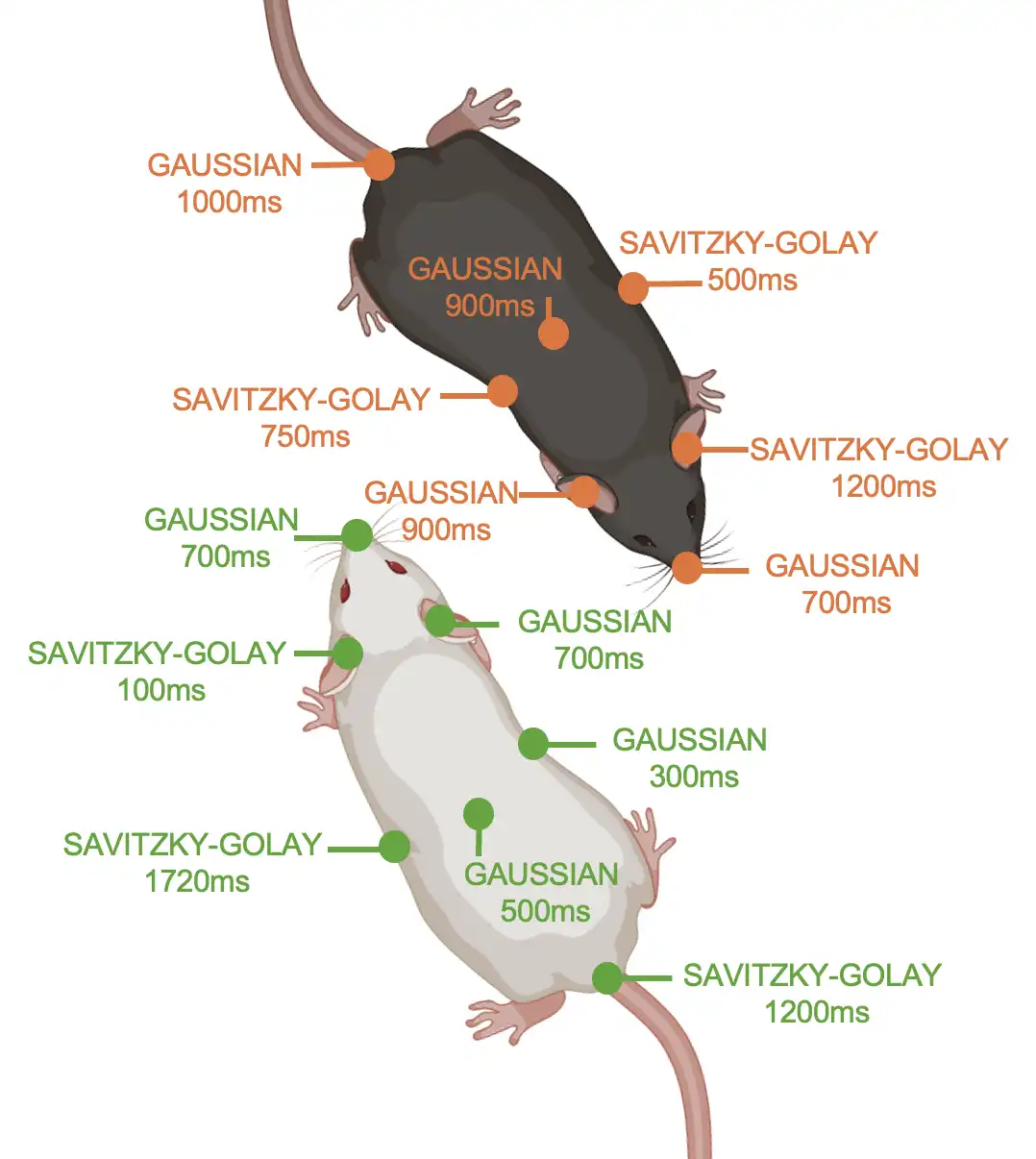
- Parameters
data_dir (str) – path to pose-estimation data in CSV or parquet format
config_path (str) – path to SimBA project config file in Configparser format.
type (Literal) – Level of smoothing: animal or body-part.
settings (Dict) – Smoothing rules for each animal or each animal body-part.
initial_import_multi_index (bool) – If True, the incoming data is multi-index columns dataframes. Use of input data is the
project_folder/csv/input_csvdirectory. Default: False.overwrite (bool) – If True, overwrites the input data. If False, then saves a copy input data in datetime-stamped sub-directory.
bool (Optional[verbose]) – If True, prints the progress. Default: True.
- Examples
>>> smoother = AdvancedSmoother(data_dir='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/csv/input_csv', >>> config_path='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', >>> type='animal', >>> settings={'Simon': {'method': 'Savitzky Golay', 'time_window': 200}, >>> 'JJ': {'method': 'Savitzky Golay', 'time_window': 200}}, >>> initial_import_multi_index=True, >>> overwrite=False) >>> smoother.run() >>> SMOOTHING_SETTINGS = {'Simon': {'Ear_left_1': {'method': 'savitzky_golay', 'time_window': 3500}, >>> 'Ear_right_1': {'method': 'gaussian', 'time_window': 500}, >>> 'Nose_1': {'method': 'savitzky_golay', 'time_window': 2000}, >>> 'Lat_left_1': {'method': 'savitzky_golay', 'time_window': 2000}, >>> 'Lat_right_1': {'method': 'gaussian', 'time_window': 2000}, >>> 'Center_1': {'method': 'savitzky_golay', 'time_window': 2000}, >>> 'Tail_base_1': {'method': 'gaussian', 'time_window': 500}}, >>> 'JJ': {'Ear_left_2': {'method': 'savitzky_golay', 'time_window': 2000}, >>> 'Ear_right_2': {'method': 'savitzky_golay', 'time_window': 500}, >>> 'Nose_2': {'method': 'gaussian', 'time_window': 3500}, >>> 'Lat_left_2': {'method': 'savitzky_golay', 'time_window': 500}, >>> 'Lat_right_2': {'method': 'gaussian', 'time_window': 3500}, >>> 'Center_2': {'method': 'gaussian', 'time_window': 2000}, >>> 'Tail_base_2': {'method': 'savitzky_golay', 'time_window': 3500}}} >>> advanced_smoother = AdvancedSmoother(config_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', >>> data_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/new_data', >>> settings=SMOOTHING_SETTINGS, type='body-part', multi_index_data=True, overwrite=False) >>> advanced_smoother.run()
Directing-other-animals calculator
- class simba.data_processors.directing_other_animals_calculator.DirectingOtherAnimalsAnalyzer(config_path: Union[str, PathLike], data_paths: Union[str, PathLike, None, List[str]] = None, bool_tables: bool = True, summary_tables: bool = False, append_bool_tables_to_features: bool = False, aggregate_statistics_tables: bool = False, verbose: bool = True, left_ear_name: Optional[str] = None, right_ear_name: Optional[str] = None, nose_name: Optional[str] = None, save_dir: Optional[Union[str, PathLike]] = None)[source]
Calculate when animals are directing towards body-parts of other animals. Results are stored in the
project_folder/logs/directionality_dataframesdirectory of the SimBA project.Note
Example expected bool table. Example expected summary table. Example expected aggregate statistics table.
Important
Requires the pose-estimation data for the
left ear,right earandnoseof each individual animals. Github Tutorial. Expected output.- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
data_paths (Optional[Union[str, os.PathLike, None]]) – Optional paths to input data files. If None, uses outlier corrected paths from project. Default: None.
bool_tables (Optional[bool]) – If True, creates boolean output tables. Default: True.
summary_tables (Optional[bool]) – If True, creates summary tables including approximate location of eye of observer and the location of observed body-parts and frames when observation was detected. Default: False.
append_bool_tables_to_features (Optional[bool]) – If True, appends boolean tables to feature data files. Default: False.
aggregate_statistics_tables (Optional[bool]) – If True, creates summary statistics tables of how much time each animal spent observing the other animals. Default: False.
left_ear_name (Optional[str]) – Name of left ear body-part. If None, SimBA will attempt to auto-detect. Default: None.
right_ear_name (Optional[str]) – Name of right ear body-part. If None, SimBA will attempt to auto-detect. Default: None.
nose_name (Optional[str]) – Name of nose body-part. If None, SimBA will attempt to auto-detect. Default: None.
- Example
>>> directing_analyzer = DirectingOtherAnimalsAnalyzer(config_path='MyProjectConfig') >>> directing_analyzer.run()
Directing-other-animals body-parts calculator
- class simba.data_processors.directing_other_animals_calculator.DirectingOtherAnimalsAnalyzer(config_path: Union[str, PathLike], data_paths: Union[str, PathLike, None, List[str]] = None, bool_tables: bool = True, summary_tables: bool = False, append_bool_tables_to_features: bool = False, aggregate_statistics_tables: bool = False, verbose: bool = True, left_ear_name: Optional[str] = None, right_ear_name: Optional[str] = None, nose_name: Optional[str] = None, save_dir: Optional[Union[str, PathLike]] = None)[source]
Calculate when animals are directing towards body-parts of other animals. Results are stored in the
project_folder/logs/directionality_dataframesdirectory of the SimBA project.Note
Example expected bool table. Example expected summary table. Example expected aggregate statistics table.
Important
Requires the pose-estimation data for the
left ear,right earandnoseof each individual animals. Github Tutorial. Expected output.- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
data_paths (Optional[Union[str, os.PathLike, None]]) – Optional paths to input data files. If None, uses outlier corrected paths from project. Default: None.
bool_tables (Optional[bool]) – If True, creates boolean output tables. Default: True.
summary_tables (Optional[bool]) – If True, creates summary tables including approximate location of eye of observer and the location of observed body-parts and frames when observation was detected. Default: False.
append_bool_tables_to_features (Optional[bool]) – If True, appends boolean tables to feature data files. Default: False.
aggregate_statistics_tables (Optional[bool]) – If True, creates summary statistics tables of how much time each animal spent observing the other animals. Default: False.
left_ear_name (Optional[str]) – Name of left ear body-part. If None, SimBA will attempt to auto-detect. Default: None.
right_ear_name (Optional[str]) – Name of right ear body-part. If None, SimBA will attempt to auto-detect. Default: None.
nose_name (Optional[str]) – Name of nose body-part. If None, SimBA will attempt to auto-detect. Default: None.
- Example
>>> directing_analyzer = DirectingOtherAnimalsAnalyzer(config_path='MyProjectConfig') >>> directing_analyzer.run()
Forward-spike-time-tiling coefficient calculator
- class simba.data_processors.fsttc_calculator.FSTTCCalculator(config_path: Union[str, PathLike], time_window: int, behavior_lst: List[str], time_delta_at_onset: Optional[bool] = False, join_bouts_within_delta: Optional[bool] = False, create_graphs: Optional[bool] = False)[source]
Compute forward spike-time tiling coefficients between pairs of classified behaviors.
- Parameters
config_path (str) – path to SimBA project config file in Configparser format.
time_window (int) – FSTTC hyperparameter; Integer representing the time window in seconds.
behavior_lst (List[str]) – Behaviors to calculate FSTTC between. FSTTC will be computed for all combinations of behaviors.
create_graphs (bool) – If True, created violin plots (as below) representing each FSTTC. Default: False.
- Parameter
Optional[bool] join_bouts_within_delta: If several bouts onsets (of the same classifier) occurs within a single time-delta, then join the bouts into a single bout.
- Parameter
Optional[bool] time_delta_at_onset: If True, time delta is initatiated at bout onset. If False, then initated at bout offset and includes bout duration. Default: False.
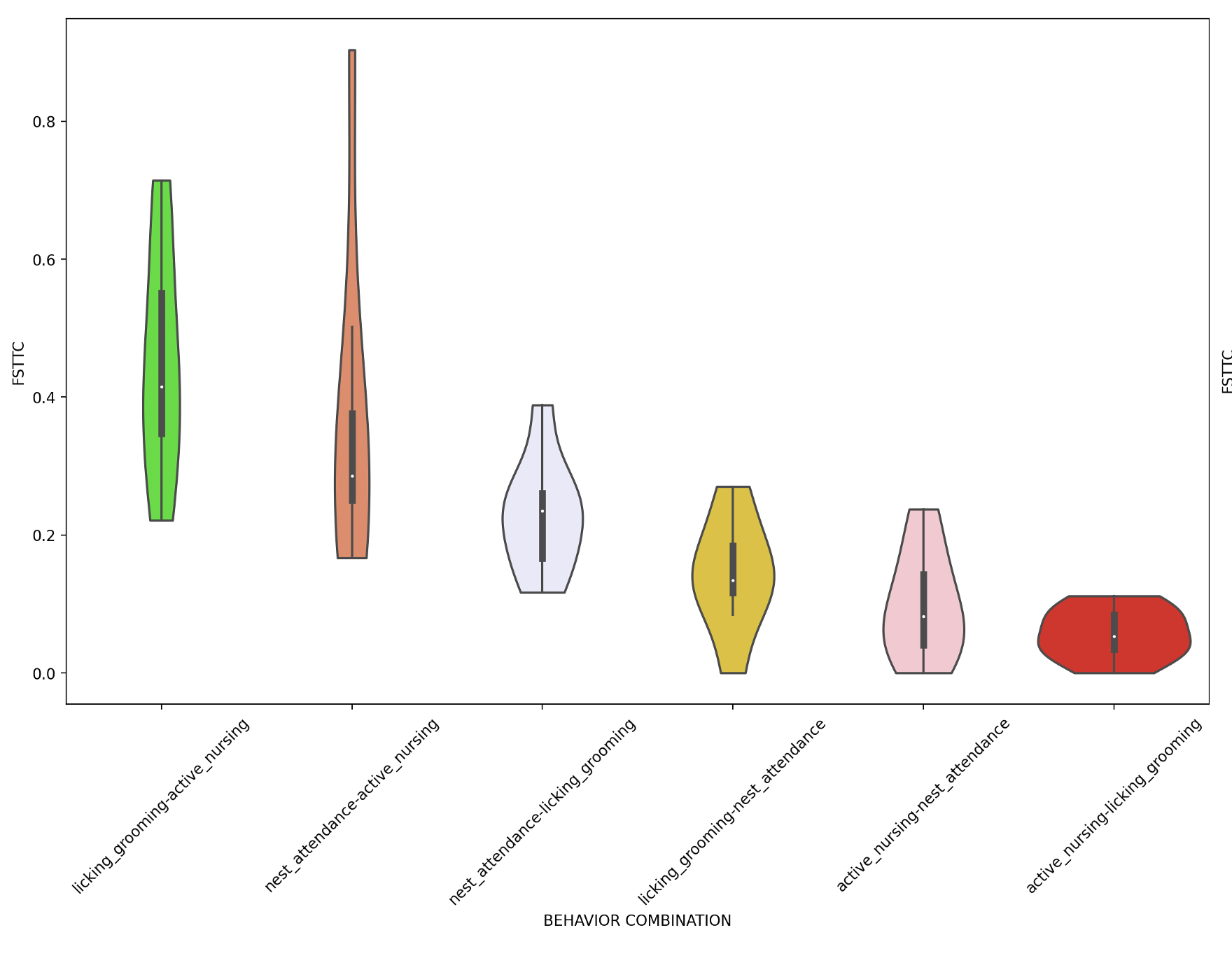
Note
Examples
>>> fsttc_calculator = FSTTCCalculator(config_path='MyConfigPath', time_window=2, behavior_lst=['Attack', 'Sniffing'], create_graphs=True) >>> fsttc_calculator.run()
References
- 1
Lee et al., Temporal microstructure of dyadic social behavior during relationship formation in mice, PLOS One, 2019.
- 2
Cutts et al., Detecting Pairwise Correlations in Spike Trains: An Objective Comparison of Methods and Application to the Study of Retinal Waves, J Neurosci, 2014.
- find_sequences()[source]
Method to create list of dataframes holding information on the sequences of behaviors including inter-temporal distances.
- Returns
Attribute – vide_df_sequence_lst
- Return type
Pose interpolator calculator
- class simba.data_processors.interpolate_pose.Interpolate(config_file_path: str, in_file: DataFrame)[source]
Interpolate missing body-parts in pose-estimation data.
- Parameters
config_file_path (str) – path to SimBA project config file in Configparser format
in_file (pd.DataFrame) – Pose-estimation data
Notes
Examples
>>> body_part_interpolator = Interpolate(config_file_path='MyProjectConfig', in_file=input_df) >>> body_part_interpolator.detect_headers() >>> body_part_interpolator.fix_missing_values(method_str='Body-parts: Nearest') >>> body_part_interpolator.reorganize_headers()
- detect_headers()[source]
Method to detect multi-index headers and set values to numeric in input dataframe
- fix_missing_values(method_str: str)[source]
Method to interpolate missing values in pose-estimation data.
- Parameters
method_str (str) – String representing interpolation method. OPTIONS: ‘None’,’Animal(s): Nearest’, ‘Animal(s): Linear’, ‘Animal(s): Quadratic’,’Body-parts: Nearest’, ‘Body-parts: Linear’, ‘Body-parts: Quadratic’
Pose interpolator and smoothing calculators
- class simba.data_processors.interpolation_smoothing.AdvancedInterpolator(data_dir: Union[str, PathLike], config_path: Union[str, PathLike], type: typing_extensions.Literal['animal', 'body-part'], settings: Dict[str, Any], initial_import_multi_index: Optional[bool] = False, overwrite: Optional[bool] = True)[source]
Interpolation method that allows different interpolation parameters for different animals or body-parts. For example, interpolate some body-parts of animals using linear interpolation, and other body-parts of animals using nearest interpolation.
- Parameters
data_dir (str) – path to pose-estimation data in CSV or parquet format
config_path (str) – path to SimBA project config file in Configparser format.
type (Literal) – Type of interpolation: animal or body-part.
settings (Dict) – Interpolation rules for each animal or each animal body-part.
initial_import_multi_index (bool) – If True, the incoming data is multi-index columns dataframes. Use of input data is the
project_folder/csv/input_csvdirectory. Default: False.overwrite (bool) – If True, overwrites the input data. If False, then saves input data in datetime-stamped sub-directory.
- Examples
>>> interpolator = AdvancedInterpolator(data_dir='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/csv/input_csv', >>> config_path='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', >>> type='animal', >>> settings={'Simon': 'linear', 'JJ': 'quadratic'}, initial_import_multi_index=True) >>> interpolator.run()
- class simba.data_processors.interpolation_smoothing.AdvancedSmoother(data_dir: Union[str, PathLike], config_path: Union[str, PathLike], type: typing_extensions.Literal['animal', 'body-part'], settings: Dict[str, Any], initial_import_multi_index: Optional[bool] = False, overwrite: Optional[bool] = True)[source]
Smoothing method that allows different smoothing parameters for different animals or body-parts. For example, smooth some body-parts of animals using Savitzky-Golay smoothing, and other body-parts of animals using Gaussian smoothing.
- Parameters
data_dir (str) – path to pose-estimation data in CSV or parquet format
config_path (str) – path to SimBA project config file in Configparser format.
type (Literal) – Level of smoothing: animal or body-part.
settings (Dict) – Smoothing rules for each animal or each animal body-part.
initial_import_multi_index (bool) – If True, the incoming data is multi-index columns dataframes. Use of input data is the
project_folder/csv/input_csvdirectory. Default: False.overwrite (bool) – If True, overwrites the input data. If False, then saves a copy input data in datetime-stamped sub-directory.
- Examples
>>> smoother = AdvancedSmoother(data_dir='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/csv/input_csv', >>> config_path='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', >>> type='animal', >>> settings={'Simon': {'method': 'Savitzky Golay', 'time_window': 200}, >>> 'JJ': {'method': 'Savitzky Golay', 'time_window': 200}}, >>> initial_import_multi_index=True, >>> overwrite=False) >>> smoother.run()
- class simba.data_processors.interpolation_smoothing.Interpolate(input_path: Union[str, PathLike], config_path: Union[str, PathLike], method: typing_extensions.Literal['Animal(s): Nearest', 'Animal(s): Linear', 'Animal(s): Quadratic', 'Body-parts: Nearest', 'Body-parts: Linear', 'Body-parts: Quadratic'], initial_import_multi_index: bool = False)[source]
Interpolate missing body-parts in pose-estimation data. “Missing” is defined as either (i) when a single body-parts is None, or when all body-parts belonging to an animal are identical (i.e., the same 2D coordinate or all None).
- Parameters
input_path (str) – Directory or file path to pose-estimation data in CSV or parquet format
config_path (str) – path to SimBA project config file in Configparser format.
str (Literal) – Type of interpolation. OPTIONS: ‘Animal(s): Nearest’, ‘Animal(s): Linear’, ‘Animal(s): Quadratic’,’Body-parts: Nearest’, ‘Body-parts: Linear’, ‘Body-parts: Quadratic’] See tutorial for info/images of the different interpolation types.
initial_import_multi_index (bool) – If True, the incoming data is multi-index columns dataframes. Default: False.

Note
Examples
>>> _ = Interpolate(input_path=data_path, config_path=SimBaProjectConfigPath, method='Animal(s): Nearest')
- class simba.data_processors.interpolation_smoothing.Smooth(config_path: str, input_path: str, time_window: int, smoothing_method: typing_extensions.Literal['Gaussian', 'Savitzky-Golay'], initial_import_multi_index: bool = False)[source]
Smooth pose-estimation data according to user-defined method.
- Parameters
input_path (str) – path to pose-estimation data in CSV or parquet format
config_path (str) – path to SimBA project config file in Configparser format.
str (Literal) – Type of smoothing_method. OPTIONS:
Gaussian,Savitzky-Golay.time_window (int) – Rolling time window in millisecond to use when smoothing. Larger time-windows and greater smoothing.
initial_import_multi_index (bool) – If True, the incoming data is multi-index columns dataframes. Default: False.
Note

- References
- Examples
>>> _ = Smooth(input_path=data_path, config_path=SimBaProjectConfigPath, smoothing_method='Savitzky-Golay', time_window=300)
Kleinberg calculator
- class simba.data_processors.kleinberg_calculator.KleinbergCalculator(config_path: Union[str, PathLike], classifier_names: Optional[List[str]] = None, sigma: float = 2, gamma: float = 0.3, hierarchy: Optional[int] = 1, verbose: bool = True, save_originals: bool = True, hierarchical_search: Optional[bool] = False, input_dir: Optional[Union[str, PathLike]] = None, output_dir: Optional[Union[str, PathLike]] = None, numba: bool = True)[source]
Smooth classification data using the Kleinberg burst detection algorithm.
Note
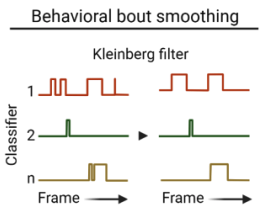
- Parameters
config_path (str) – path to SimBA project config file in Configparser format
classifier_names (List[str]) – Classifier names to apply Kleinberg smoothing to.
sigma (float) – State transition cost for moving to higher burst levels. Higher values (e.g., 2-3) produce fewer but longer bursts; lower values (e.g., 1.1-1.5) detect more frequent, shorter bursts. Must be > 1.01. Default: 2.
gamma (float) – State transition cost for moving to lower burst levels. Higher values (e.g., 0.5-1.0) reduce total burst count by making downward transitions costly; lower values (e.g., 0.1-0.3) allow more flexible state changes. Must be >= 0. Default: 0.3.
hierarchy (int) – Hierarchy level to extract bursts from (0=lowest, higher=more selective). Level 0 captures all bursts; level 1-2 typically filters noise; level 3+ selects only the most prominent, sustained bursts. Higher levels yield fewer but more confident detections. Must be >= 0. Default: 1.
hierarchical_search (bool) – If True, searches for target hierarchy level within detected burst periods, falling back to lower levels if target not found. If False, extracts only bursts at the exact specified hierarchy level. Recommended when target hierarchy may be sparse. Default: False.
input_dir (Optional[Union[str, os.PathLike]]) – The directory with files to perform kleinberg smoothing on. If None, defaults to project_folder/csv/machine_results
output_dir (Optional[Union[str, os.PathLike]]) – Location to save smoothened data in. If None, defaults to project_folder/csv/machine_results
save_originals (Optional[bool]) – If True, saves the original data in sub-directory of the ouput directory.`
numba (bool) – If True, uses Numba JIT-accelerated burst detection for faster computation. Default: False.
- Example I
>>> kleinberg_calculator = KleinbergCalculator(config_path='MySimBAConfigPath', classifier_names=['Attack'], sigma=2, gamma=0.3, hierarchy=2, hierarchical_search=False) >>> kleinberg_calculator.run()
- Example 2
>>> output_dir = r'/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/csv/kleinberg_gridsearch_test' >>> input_dir = r'/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/csv/kleinberg_gridsearch_test' >>> kleinberg_calculator = KleinbergCalculator(config_path=r'/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', classifier_names=['Attack', 'Sniffing', 'Rear'], sigma=2, gamma=0.3, hierarchy=3, hierarchical_search=False, input_dir=input_dir, output_dir=output_dir)
References
- 1
Kleinberg, Bursty and Hierarchical Structure in Streams, Data Mining and Knowledge Discovery, vol. 7, pp. 373–397, 2003.
- 2
Lee et al., Temporal microstructure of dyadic social behavior during relationship formation in mice, PLOS One, 2019.
- 3
Bordes et al., Automatically annotated motion tracking identifies a distinct social behavioral profile following chronic social defeat stress, bioRxiv, 2022.
- 4
Chanthongdee et al., Comprehensive ethological analysis of fear expression in rats using DeepLabCut and SimBA machine learning model. Front. Behav. Neurosci. https://doi.org/10.3389/fnbeh.2024.1440601
Movement calculator
- class simba.data_processors.movement_calculator.MovementCalculator(config_path: Union[str, PathLike], body_parts: Union[List[str], Tuple[str]], threshold: float = 0.0, file_paths: Optional[List[str]] = None, save_path: Optional[Union[str, PathLike]] = None, distance: bool = True, velocity: bool = True, video_time_stamps: Optional[Dict[str, tuple]] = None, transpose: bool = False, verbose: bool = True, frame_count: bool = False, video_length: bool = False)[source]
Compute aggregate movement statistics from pose-estimation data in SimBA project.
- Parameters
config_path (Union[str, os.PathLike]) – path to SimBA project config file in Configparser format
body_parts (List[str]) – Body-parts to use for movement calculations OR
Animal_name CENTER OF GRAVITY. IfAnimal_name CENTER OF GRAVITY, then SimBA will approximate animal centroids through convex hull.threshold (float) – Filter body-part detection below set threshold (Value 0-1). Default: 0.00
file_paths (Optional[List[str]]) – Files to calculate movements for. If None, then all files in
project_folder/csv/outlier_corrected_movement_locationdirectory.save_path (Optional[Union[str, os.PathLike]]) – Path to save the movement log. If None, saves to
project_folder/logs/Movement_log_{datetime}.csv. Default: Nonedistance (bool) – If True, calculate distance metrics. Default: True
velocity (bool) – If True, calculate velocity metrics. Default: True
video_time_stamps (Optional[Dict[str, dict]]) – Dictionary mapping video file names (without extension) to time windows. Each value is a dict with
STARTandENDkeys (in seconds). Only frames within the time window are analyzed. If None, all frames are used. Default: None.transpose (bool) – If True, transpose the results DataFrame. Default: False
verbose (bool) – If True, print progress messages. Default: True
frame_count (bool) – If True, include frame count in results. Default: False
video_length (bool) – If True, include video length in results. Default: False
Note
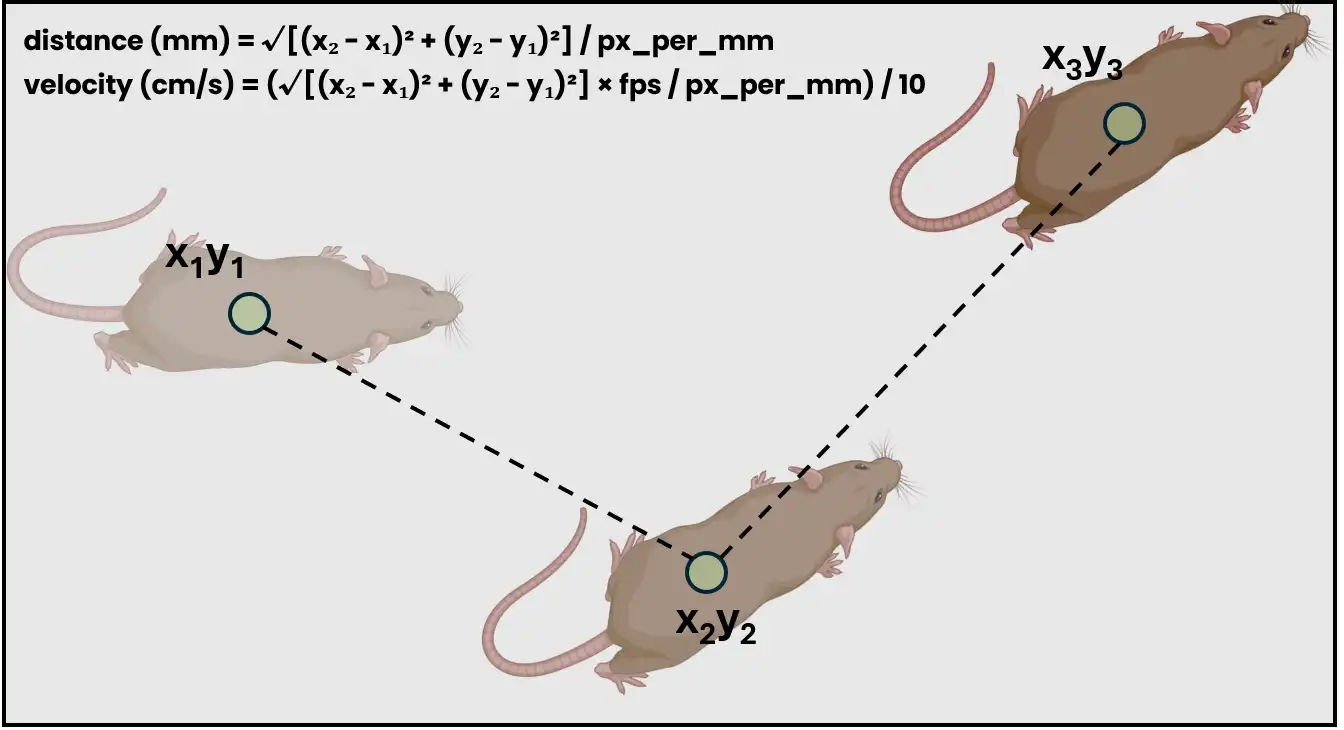
- Examples
>>> body_parts=['Animal_1 CENTER OF GRAVITY'] >>> movement_processor = MovementCalculator(config_path='project_folder/project_config.ini', body_parts=body_parts) >>> movement_processor.run() >>> movement_processor.save() >>> time_stamps = pd.read_csv('mastersheet.csv')[['VIDEO_FILE_NAME', 'START', 'END']].set_index('VIDEO_FILE_NAME').to_dict(orient='index') >>> movement_processor = MovementCalculator(config_path='project_folder/project_config.ini', body_parts=['center'], video_time_stamps=time_stamps) >>> movement_processor.run() >>> movement_processor.save()
Pup-retrieval calculator
- class simba.data_processors.pup_retrieval_calculator.PupRetrieverCalculator(config_path: str, settings: Dict[str, Union[float, str, bool]])[source]
Pup retreival calculator used in
Winters et al., `Sci Reports`, 2022- Parameters
Note

- Examples
>>> settings = {'pup_track_p': 0.025, 'dam_track_p': 0.5, 'start_distance_criterion': 80.0, 'carry_frames': 90.0, 'core_nest': 'corenest', 'nest': 'nest', 'dam_name': '1_mother', 'pup_name': '2_pup', 'smooth_function': 'gaussian', 'smooth_factor': 5, 'max_time': 90.0, 'clf_carry': 'carry', 'clf_approach': 'approach', 'clf_dig': 'digging', 'distance_plots': True, 'log': True, 'swarm_plot': True} >>> config_path = '/Users/simon/Downloads/Automated PRT_test/project_folder/project_config.ini' >>> calculator = PupRetrieverCalculator(config_path=config_path, settings=settings) >>> calculator.run()
- References
- 1
Winters, Carmen, Wim Gorssen, Victoria A. Ossorio-Salazar, Simon Nilsson, Sam Golden, and Rudi D’Hooge. “Automated Procedure to Assess Pup Retrieval in Laboratory Mice.” Scientific Reports 12, no. 1 (2022): 1663. https://doi.org/10.1038/s41598-022-05641-w.
SimBA pyburst calculator
- simba.data_processors.pybursts_calculator.kleinberg_burst_detection(offsets: ndarray, s: float, gamma: float)[source]
Detect hierarchical bursts in a 1D sequence of event times using Kleinberg’s two-state infinite-state automaton (modified from pybursts).
Bursts are intervals where events arrive at a higher-than-baseline rate. The algorithm assigns each inter-event gap to a discrete level
q: level0is baseline, higher levels (1, 2, …) are progressively faster (higher-rate) bursts. Each level transition opens or closes a burst at that level, producing a hierarchy of nested bursts.Note
Private helper used by
KleinbergCalculator. For an end-to-end pipeline (frame indices → bouts → bursts), use that class.- Parameters
offsets (np.ndarray) – 1D numeric array of event times (seconds, frame indices, or any monotonically meaningful unit). May be unsorted; sorted internally. Must have strictly positive gaps (no two events at the same time).
s (float) – Base of the rate scale (
> 1). The candidate rate at leveljiss**j / mean_gap, so largersmeans levels grow farther apart and bursts must be more pronounced to reach higher levels. Common choice:s = 2.gamma (float) – Cost of moving up one level (
>= 0). Highergammapenalizes rising into a burst, producing fewer / shorter bursts. Lowergammamakes the detector more sensitive. Common choice:gamma = 1.
- Returns
2D
np.ndarrayof shape(N, 3)anddtype=objectwith one row per detected burst, columns[level, start_offset, end_offset]:level— integer burst level (0is the baseline level, higher levels are nested faster bursts).start_offset— value fromoffsetswhere this burst opens.end_offset— value fromoffsetswhere this burst closes (inclusive of the last event in the run).
For a single-event input, returns a single row
[0, offsets[0], offsets[0]].- Return type
np.ndarray
- Raises
ValueError – If
offsetscontains two or more events at the same time (zero gap).
See also
kleinberg_burst_detection()for Numba JIT-accelerated version.EXPECTED RUNTIMES pyburst numbaN
BURSTS
NON-NUMBA TIME (S)
NUMBA TIME (S)
SPEEDUP
100
7
0.0223
0.0001
~223x
1,000
25
0.2751
0.0006
~458x
10,000
81
7.4712
0.0098
~762x
100,000
390
443.0849
0.1170
~3,787x
1,000,000
2,020
N/A
1.6230
N/A
S: 2.0
GAMMA: 0.3
GAPS: exponential (scale=1.0)
- Example
>>> import numpy as np >>> from simba.data_processors.pybursts_calculator import kleinberg_burst_detection >>> offsets = np.array([1.0, 1.1, 1.2, 5.0, 9.0, 9.05, 9.1]) >>> bursts = kleinberg_burst_detection(offsets=offsets, s=2.0, gamma=1.0) >>> bursts.shape[1] 3
Severity calculator
- class simba.data_processors.severity_calculator.SeverityCalculator(config_path: Union[str, PathLike], settings: Dict)[source]
Computes the “severity” of classification frame events based on how much the animals are moving. Frames are scored as less or more severe at lower and higher movements, respectively.
- Parameters
Note
Examples
>>> settings = {'brackets': 10, 'clf': 'Attack', 'animals': ['Simon', 'JJ'], 'time': True, 'frames': False} >>> processor = SeverityCalculator(config_path='project_folder/project_config.ini', settings=settings) >>> processor.run() >>> processor.save()
Classifier time-bins calculator
- class simba.data_processors.timebins_clf_calculator.TimeBinsClfCalculator(config_path: Union[str, PathLike], bin_length: int, classifiers: List[str], data_path: Optional[Union[str, PathLike]] = None, first_occurrence: bool = False, event_count: bool = False, total_event_duration: bool = True, mean_event_duration: bool = False, median_event_duration: bool = False, mean_interval_duration: bool = False, median_interval_duration: bool = False, include_timestamp: bool = False, transpose: bool = False)[source]
Computes aggregate classification results in user-defined time-bins. Results are stored in the
project_folder/logsdirectory of the SimBA project.- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
bin_length (int) – Integer representing the time bin size in seconds.
classifiers (List[str]) – Names of classifiers to calculate aggregate statistics in time-bins for. EXAMPLE: [‘Attack’, ‘Sniffing’]
data_path (Optional[Union[str, os.PathLike]]) – Optional path to directory containing CSV files or single CSV file. If None, uses machine results from project. Default: None.
first_occurrence (bool) – If True, calculate first occurrence time for each classifier in each time bin. Default: False.
event_count (bool) – If True, calculate event count for each classifier in each time bin. Default: False.
total_event_duration (bool) – If True, calculate total event duration for each classifier in each time bin. Default: True.
mean_event_duration (bool) – If True, calculate mean event duration for each classifier in each time bin. Default: False.
median_event_duration (bool) – If True, calculate median event duration for each classifier in each time bin. Default: False.
mean_interval_duration (bool) – If True, calculate mean interval duration between events for each classifier in each time bin. Default: False.
median_interval_duration (bool) – If True, calculate median interval duration between events for each classifier in each time bin. Default: False.
include_timestamp (bool) – If True, include START TIME and END TIME (in HH:MM:SS format) columns in output. Default: False.
transpose (bool) – If True, transpose results with MultiIndex columns (CLASSIFIER, TIME BIN #, MEASUREMENT) so one video per row Default: False.
Note
- Example
>>> timebin_clf_analyzer = TimeBinsClfCalculator(config_path='MyConfigPath', bin_length=15, classifiers=['Attack', 'Sniffing'], event_count=True, total_event_duration=True) >>> timebin_clf_analyzer.run() >>> timebin_clf_analyzer.save()
Movement time-bins calculator
- class simba.data_processors.timebins_movement_calculator.TimeBinsMovementCalculator(config_path: Union[str, PathLike], bin_length: Union[int, float], body_parts: Union[List[str], Tuple[str]], data_path: Optional[Union[List[Union[str, PathLike]], str, PathLike]] = None, plots: bool = False, verbose: bool = True, threshold: float = 0.0, distance: bool = True, velocity: bool = True, transpose: bool = False, include_timestamp: bool = False)[source]
Compute aggregate movement and/or velocity statistics in user-defined time-bins.
Note

See also
For multicore processing, see
simba.data_processors.timebins_movement_calculator_mp.TimeBinsMovementCalculator.- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file.
body_parts (Union[List[str], Tuple[str]]) – Body-part names to include in the movement calculations.
data_path (Optional[Union[List[Union[str, os.PathLike]], Union[str, os.PathLike]]]) – Optional file path(s) to process. If
None, all outlier-corrected files in the project are used.plots (bool) – If
True, create per-video movement line plots for each body-part. Default:False.(bool) (verbose) – If True, prints progress messages during processing. Default: True.
threshold (float) – Confidence threshold used when filtering low-confidence positions. Default:
0.0.distance (bool) – If
True, compute movement distance per time-bin. Default:True.velocity (bool) – If
True, compute velocity per time-bin. Default:True.transpose (bool) – If
True, save output in transposed format with one column per time-bin. Default:False.include_timestamp (bool) – If
True, include start/end timestamps for each time-bin in saved results. Default:False.
- Example
>>> calculator = TimeBinsMovementCalculator(config_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', bin_length=0.04, plots=True, body_parts=['Nose_1', 'Nose_2']) >>> calculator.run()
Movement time-bins calculator (multiprocess)
- class simba.data_processors.timebins_movement_calculator_mp.TimeBinsMovementCalculatorMultiprocess(config_path: Union[str, PathLike], bin_length: Union[int, float], body_parts: Union[List[str], Tuple[str]], data_path: Optional[Union[List[Union[str, PathLike]], str, PathLike]] = None, plots: bool = False, verbose: bool = True, core_cnt: int = -1, distance: bool = True, velocity: bool = True, threshold: float = 0.0, transpose: bool = False, include_timestamp: bool = False)[source]
Computes aggregate movement statistics in user-defined time-bins using multiprocessing for improved performance.
Note
Note
On macOS (Darwin), multiprocessing start method is automatically set to ‘spawn’ for compatibility.

See also
For single core class, see
simba.data_processors.timebins_movement_calculator.TimeBinsMovementCalculator().- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
bin_length (Union[int, float]) – Time bin size in seconds. Must be greater than 0.
body_parts (List[str]) – List of body part names to calculate movement for (e.g., [‘Nose_1’, ‘Nose_2’]). Body parts must exist in the project’s body part configuration.
data_path (Optional[List[Union[str, os.PathLike]]]) – Optional list of specific file paths to process. If None, processes all files in the project’s outlier corrected directory. Can also be a single file path or a directory path containing CSV files.
plots (Optional[bool]) – If True, creates time-bin line plots representing the movement in each time-bin per video. Results are saved in the
project_folder/logs/sub-directory. Default: False.verbose (bool) – If True, prints progress information during processing. Default: True.
core_cnt (int) – Number of CPU cores to use for multiprocessing. If -1, uses all available cores. If greater than available cores, uses all available cores. Must be greater than 0. Default: -1.
- Example
>>> calculator = TimeBinsMovementCalculatorMultiprocess( ... config_path='/Users/simon/Desktop/envs/simba/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', ... bin_length=5.0, ... body_parts=['Nose_1', 'Nose_2'], ... plots=True, ... core_cnt=4 ... ) >>> calculator.run() >>> calculator.save()
Aggregate classifier statistics (multiprocess)
- class simba.data_processors.agg_clf_counter_mp.AggregateClfCalculatorMultiprocess(config_path: Union[str, PathLike], classifiers: List[str], data_dir: Optional[Union[str, PathLike]] = None, detailed_bout_data: bool = False, transpose: bool = False, first_occurrence: bool = True, event_count: bool = True, total_event_duration: bool = True, mean_event_duration: bool = True, median_event_duration: bool = True, mean_interval_duration: bool = True, median_interval_duration: bool = True, pct_of_session: bool = True, frame_count: bool = False, video_length: bool = False, verbose: bool = True, core_cnt: int = -1)[source]
Compute aggregate descriptive statistics from classification data using multiprocessing.
This class analyzes machine learning classifier results to calculate various descriptive statistics such as bout counts, durations, intervals, and first occurrences for each classifier in each video. Results can be saved in detailed or summary formats.
See also
For single core class, see
simba.data_processors.agg_clf_calculator.AggregateClfCalculator()- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file in Configparser format.
classifiers (List[str]) – List of classifier names to calculate aggregate statistics for. Must be valid classifier names from the project.
data_dir (Optional[Union[str, os.PathLike]]) – Directory containing the machine results CSV files. If None, uses
project_folder/csv/machine_results.detailed_bout_data (bool) – If True, saves detailed bout data (start frame, end frame, bout time, etc.) for each bout in each video. Default: False.
transpose (bool) – If True, creates output with one video per row. If False, one measurement per row. Default: False.
first_occurrence (bool) – If True, calculates first occurrence time for each classifier. Default: True.
event_count (bool) – If True, calculates total number of bouts for each classifier. Default: True.
total_event_duration (bool) – If True, calculates total duration of all bouts for each classifier. Default: True.
mean_event_duration (bool) – If True, calculates mean duration of bouts for each classifier. Default: True.
median_event_duration (bool) – If True, calculates median duration of bouts for each classifier. Default: True.
mean_interval_duration (bool) – If True, calculates mean interval between bouts for each classifier. Default: True.
median_interval_duration (bool) – If True, calculates median interval between bouts for each classifier. Default: True.
frame_count (bool) – If True, includes total frame count in the output. Default: False.
video_length (bool) – If True, includes video length in seconds in the output. Default: False.
core_cnt (int) – Number of CPU cores. If -1, then all available cores.
- Example
>>> clf_calculator = AggregateClfCalculatorMultiprocess( ... config_path="project_folder/project_config.ini", ... classifiers=['Attack', 'Sniffing'], ... detailed_bout_data=True, ... transpose=True ... ) >>> clf_calculator.run() >>> clf_calculator.save()
Directing-animal-to-bodypart calculator
- class simba.data_processors.directing_animal_to_bodypart.DirectingAnimalsToBodyPartAnalyzer(config_path: Union[str, PathLike])[source]
Calculate when animals are directing towards their own body-parts. Results are stored in the
project_folder/logs/directionality_dataframesdirectory of the SimBA project.- Parameters
config_path (Union[str, os.PathLike]) – path to SimBA project config file in Configparser format
Important
Requires the pose-estimation data for the
left ear,right earandnoseof each individual animals. Tutorial. Expected output.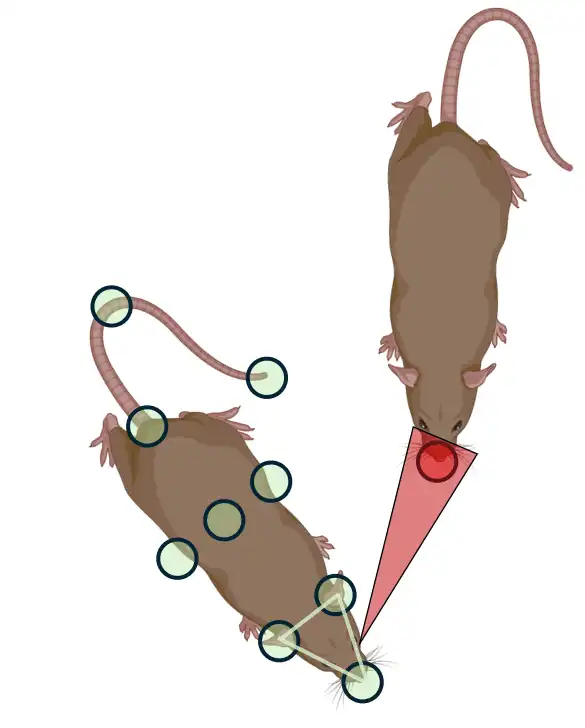
- Example
>>> directing_analyzer = DirectingAnimalsToBodyPartAnalyzer(config_path='MyProjectConfig') >>> directing_analyzer.process_directionality() >>> directing_analyzer.create_directionality_dfs() >>> directing_analyzer.save_directionality_dfs() >>> directing_analyzer.summary_statistics()
- create_directionality_dfs()[source]
Method to transpose results created by
process_directionality(). into dict of dataframes- Returns
Attribute – directionality_df_dict
- Return type
- process_directionality()[source]
Method to compute when animals are directing towards their own body-parts.
- Returns
Attribute – results_dict
- Return type
Severity bout-based calculator
- class simba.data_processors.severity_bout_based_calculator.SeverityBoutCalculator(config_path: Union[str, PathLike], settings: Dict)[source]
Computes the “severity” of classification bout events based on how much the animals are moving within the bout. Bouts are scored as less or more severe at lower and higher movements, respectively.
- Parameters
config_path (str) – path to SimBA project config file in Configparser format.
settings (dict) – how to calculate the severity. E.g., {‘brackets’: 10, ‘clf’: ‘Attack’, ‘animals’: [‘Simon’, ‘JJ’], ‘normalization’: ‘ALL VIDEOS’, ‘save_bin_definitions’: True, ‘visualize’: True, ‘visualize_event_cnt’: ‘ALL’, ‘video_speed’: 1.0, ‘show_pose’: True}
Note
Examples
>>> settings = {'brackets': 10, 'clf': 'Attack', 'animals': ['Simon', 'JJ'], 'normalization': 'ALL VIDEOS', 'save_bin_definitions': True, 'visualize': True, 'visualize_event_cnt': 'ALL', 'video_speed': 1.0, 'show_pose': True} >>> processor = SeverityBoutCalculator(config_path='project_folder/project_config.ini', settings=settings) >>> processor.run() >>> processor.save()
Severity frame-based calculator
- class simba.data_processors.severity_frame_based_calculator.SeverityFrameCalculator(config_path: Union[str, PathLike], settings: Dict)[source]
Computes the “severity” of classification frame events based on how much the animals are moving. Frames are scored as less or more severe at lower and higher movements, respectively.
- Parameters
Note
Examples
>>> settings = {'brackets': 10, 'clf': 'Attack', 'animals': ['Simon', 'JJ'], 'time': True, 'frames': False, 'normalization': 'ALL VIDEOS', 'save_bin_definitions': True} >>> processor = SeverityFrameCalculator(config_path='project_folder/project_config.ini', settings=settings) >>> processor.run() >>> processor.save()
YOLO track cleaner
SHAP log (GPU)
- simba.data_processors.cuda.create_shap_log.create_shap_log(rf_clf: Union[str, PathLike, RandomForestClassifier], x: Union[DataFrame, ndarray], y: Union[DataFrame, Series, ndarray], cnt_present: int, cnt_absent: int, x_names: Optional[List[str]] = None, clf_name: Optional[str] = None, save_dir: Optional[Union[str, PathLike]] = None, verbose: Optional[bool] = True) Union[None, Tuple[DataFrame, DataFrame, int]][source]
Computes SHAP (SHapley Additive exPlanations) values using a GPU for a RandomForestClassifier, based on specified counts of positive and negative samples, and optionally saves the results.
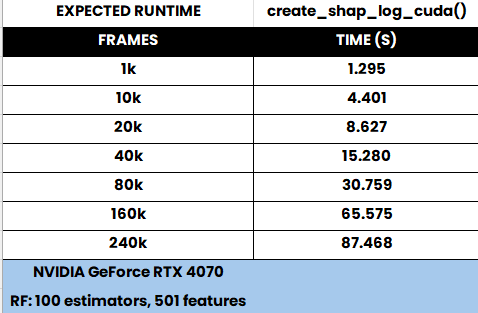
Note
The SHAP library has to be built from git repo rather than pip: pip install git+https://github.com/slundberg/shap.git.
The scikit model cannot be built using max_depth > 31. You can set this in the SimBA config under [create ensemble settings][rf_max_depth], or rf_max_depth in the config CSV’s.
- Parameters
rf_clf (Union[str, os.PathLike, RandomForestClassifier]) – Trained RandomForestClassifier model or path to the saved model. Can be a string, os.PathLike object, or an instance of RandomForestClassifier.
x (Union[pd.DataFrame, np.ndarray]) – Input features used for SHAP value computation. Can be a pandas DataFrame or numpy ndarray.
y (Union[pd.DataFrame, pd.Series, np.ndarray]) – Target labels corresponding to the input features. Can be a pandas DataFrame, pandas Series, or numpy ndarray with 0 and 1 values.
cnt_present (int) – Number of positive samples (label=1) to include in the SHAP value computation.
cnt_absent (int) – Number of negative samples (label=0) to include in the SHAP value computation.
x_names (Optional[List[str]]) – Optional list of feature names corresponding to the columns in x. If x is a DataFrame, this is extracted automatically.
clf_name (Optional[str]) – Optional name for the classifier, used in naming output files. If not provided, it is extracted from the y labels if possible.
save_dir (Optional[Union[str, os.PathLike]]) – Optional directory path where the SHAP values and corresponding raw features are saved as CSV files.
verbose (Optional[bool]) – Optional boolean flag indicating whether to print progress messages. Defaults to True.
- Return Union[None, Tuple[pd.DataFrame, pd.DataFrame, int]]
If save_dir is None, returns a tuple containing: - V: DataFrame with SHAP values, expected value, sum of SHAP values, prediction probability, and target labels. - R: DataFrame containing the raw feature values for the selected samples. - expected_value: The expected value from the SHAP explainer.
If save_dir is provided, the function returns None and saves the output to CSV files in the specified directory.
- Example
>>> x = np.random.random((1000, 501)).astype(np.float32) >>> y = np.random.randint(0, 2, size=(len(x), 1)).astype(np.int32) >>> clf_names = [str(x) for x in range(501)] >>> results = create_shap_log(rf_clf=MODEL_PATH, x=x, y=y, cnt_present=int(i/2), cnt_absent=int(i/2), clf_name='TEST', x_names=clf_names, verbose=False)
Mutual exclusivity refactorer
- class simba.data_processors.mutual_exclusivity_corrector.MutualExclusivityCorrector(rules: dict, config_path: Union[str, PathLike])[source]
Refactor classification results according to user-defined mutual exclusivity rules.
Note
- Examples
>>> rules = {1: {'rule_type': 'threshold_determinator','determinator': 'Attack', 'threshold': 0.5, 'subordinates': ['Sniffing']}, 2: {'rule_type': 'threshold_determinator', 'determinator': 'Attack', 'threshold': 0.0, 'subordinates': ['Sniffing', 'Rear']}} >>> exclusivity_corrector = MutualExclusivityCorrector(config_path='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', rules=rules) >>> exclusivity_corrector.run() >>> rules = {1: {'rule_type': 'highest_probability', 'subordinates': ['body', 'face'], 'winner': 'body', 'skip_files_with_identical': True}} >>> exclusivity_corrector = MutualExclusivityCorrector(config_path='/Users/simon/Desktop/envs/troubleshooting/two_black_animals_14bp/project_folder/project_config.ini', rules=rules) >>> exclusivity_corrector.run()
Boolean conditional calculator
- class simba.data_processors.boolean_conditional_calculator.BooleanConditionalCalculator(config_path: Union[str, PathLike], rules: Dict[str, Union[bool, str]], data_path: Optional[Union[str, PathLike]] = None, agg_save_path: Optional[Union[str, PathLike]] = None, detailed_save_path: Optional[Union[str, PathLike]] = None, verbose: bool = True)[source]
Compute descriptive statistics (e.g., the time in seconds and number of frames) of multiple Boolean fields fullfilling user-defined conditions.
For example, computedescriptive statistics for when Animal 1 is inside the shape Rectangle_1 while at the same time directing towards shape Polygon_1, while at the same time Animal 2 is outside shape Rectangle_1 and directing towards Polygon_1.
- param Union[str, os.PathLike] config_path
path to SimBA project config file in Configparser format.
- param Dict[str, Union[bool, str]] rules
Rules with field names as keys and bools (or string representations of bools) as values.
- param Optional[Union[str, os.PathLike, None]] data_path
Optional data paths to be processsed. Can be a directory or file path. If None, all CSVs inside the projecet_folder/csv/outlier_corrected_movement_location are analysed.
- param Optional[Union[str, os.PathLike]] agg_save_path
Optional location where to save the aggregate results as CSV file. If None, then results are saved in project logs folder under the
Detailed_conditional_aggregate_statistics_{self.datetime}.csvfilename.- param Optional[Union[str, os.PathLike]] detailed_save_path
Optional location where to save the detailed results as CSV file (bout level data). If None, then results are saved in project logs folder under the
Detailed_conditional_aggregate_statistics_{self.datetime}.csvfilename.
- example I
>>> rules = {'Rectangle_1 Simon in zone': 'TRUE', 'Polygon_1 JJ in zone': 'TRUE'} # OR {'Rectangle_1 Simon in zone': True, 'Polygon_1 JJ in zone': True} >>> conditional_bool_rule_calculator = BooleanConditionalCalculator(rules=rules, config_path='/Users/simon/Desktop/envs/troubleshooting/two_animals_16bp_032023/project_folder/project_config.ini') >>> conditional_bool_rule_calculator.run() >>> conditional_bool_rule_calculator.save()
- example II
>>> rules = {'Stimulus 2 Animal_1 in zone': True, 'Stimulus 6 Animal_1 in zone': 'falsE'} >>> runner = BooleanConditionalCalculator(rules=rules, config_path=r"C: roubleshooting\RAT_NOR\project_folder\project_config.ini", data_path=r'C: roubleshooting\RAT_NOR\project_folder\csv
- eatures_extracted’)
>>> runner.run() >>> runner.save()
- references
- 1
Shonka, Sophia, and Michael J Hylin. “Younger Is Better But Only for Males: Social Behavioral Development Following Juvenile Traumatic Brain Injury to the Prefrontal Cortex,” n.d.
Gibbs sampling
- class simba.data_processors.gibbs_sampler.GibbSampler(data: ndarray, save_path: Union[str, PathLike], sequence_length: int = 4, iterations: int = 1500, epochs: int = 2, stop_val: float = 0.001, pseudo_number: float = 1e-05, plateau_val: int = 50)[source]
Gibbs sampling for finding “motifs” in categorical sequences.
- Parameters
data (np.ndarray) – 2-dimensional array where observations are organised by row and each sequential sample in the observation is organized by column.
save_path (Union[str, os.PathLike]) – The path location where to save the CSV results.
sequence_length (int) – The length of the motif sequence searched for.
iterations (int) – Number of iterations per epoch. Default: 1500.
epochs (int) – Number of epochs of iterations. Default: 4.
stop_val (float) – Terminate once the error value reaches below this threshold. Default 0.001.
plateau_val (int) – Terminate epoch when error rate has remained unchanged for
plateau_valcount of iterations. Default 50.pseudo_number (float) – Small error value for fuzzy search and avoid division by zero errors. Default: 10e-6.
- Example
>>> data = pd.read_csv(r"/Users/simon/Desktop/envs/simba/simba/tests/data/sample_data/gibbs_sample_cardinal.csv", index_col=0).values >>> sampler = GibbSampler(data=data, save_path=r'/Users/simon/Desktop/gibbs.csv', epochs=5, iterations=600) >>> sampler.run()
- References
- 1
Lawrence et al, Detecting Subtle Sequence Signals: a Gibbs Sampling Strategy for Multiple Alignment, Science, vol. 262, pp. 208-214, 1993.
- 2
Great YouTube tutorial / explanation by Xiaole Shirley Liu - https://www.youtube.com/watch?v=NRjhfyXWHuQ.
- 3
Weinreb et al. Keypoint-MoSeq: parsing behavior by linking point tracking to pose dynamics, Nature Methods, 21, 1329–1339 (2024).
Spontaneous alternation calculator
- class simba.data_processors.spontaneous_alternation_calculator.SpontaneousAlternationCalculator(config_path: Union[str, PathLike], arm_names: List[str], center_name: str, animal_area: Optional[int] = 80, threshold: Optional[float] = 0.0, buffer: Optional[int] = 2, verbose: Optional[bool] = False, detailed_data: Optional[bool] = False, data_path: Optional[Union[str, PathLike]] = None)[source]
Compute spontaneous alternations based on specified ROIs and animal detection parameters.
Note
This method computes spontaneous alternation by fitting the smallest viable geometry (a.k.a. shape, polygon) that encompasses the animal key-points (but see buffer parameter below). It then checks the percent overlap between the animal geometry and the defined arm and center geometries in each frame. If the percent overlap is more or equal to the specified threshold, then the animal considered to visiting the relevant arm. The animal is considered exiting an arm when the percent overlap with a different ROI is above the threshold.
Attention
Requires SimBA project with (i) only one tracked animal, (ii) at least three pose-estmated body-parts, and (iii) defined ROIs representing the arms and the center of the maze.
- Parameters
config_path (Union[str, os.PathLike]) – Path to SimBA project config file.
arm_names (List[str]) – List of ROI names representing the arms.
center_name (str) – Name of the ROI representing the center of the maze
animal_area (Optional[int]) – Value between 51 and 100, representing the percent of the animal body that has to be situated in a ROI for it to be considered an entry.
threshold (Optional[float]) – Value between 0.0 and 1.0. Body-parts with detection probabilities below this value will be (if possible) filtered when constructing the animal geometry.
buffer (Optional[int]) – Millimeters area for which the animal geometry should be increased in size. Useful if the animal geometry does not fully cover the animal.
detailed_data (Optional[bool]) – If True, saves an additional CSV for each analyzed video with detailed data pertaining each error type and alternation sequence.
data_path (Optional[Union[str, os.PathLike]]) – Directory of path to the file to be analyzed. If None, then
project_folder/outlier_corrected_movement_locationdirectory.
- Example
>>> x = SpontaneousAlternationCalculator(config_path='/Users/simon/Desktop/envs/simba/troubleshooting/spontenous_alternation/project_folder/project_config.ini', arm_names=['A', 'B', 'C'], center_name='Center', threshold=0.0, animal_area=100, buffer=2, detailed_data=True) >>> x.run() >>> x.save()
“Blob” location detector
- class simba.data_processors.blob_location_computer.BlobLocationComputer(data_path: Union[str, PathLike], verbose: Optional[bool] = True, gpu: Optional[bool] = True, batch_size: int = 2500, save_dir: Optional[Union[str, PathLike]] = None, smoothing: Optional[str] = None, multiprocessing: bool = False)[source]
Detecting and saving blob locations from video files.
- Parameters
data_path (Union[str, os.PathLike]) – Path to a video file or a directory containing video files. The videos will be processed for blob detection.
verbose (Optional[bool]) – If True, prints progress and success messages to the console. Default is True.
gpu (Optional[bool]) – If True, GPU acceleration will be used for blob detection. Default is True.
batch_size (Optional[int]) – The number of frames to process in each batch for blob detection. Default is 2500.
save_dir (Optional[Union[str, os.PathLike]]) – Directory where the blob location data will be saved as CSV files. If None, the results will not be saved. Default is None.
multiprocessing (Optional[bool]) – If True, video background subtraction will be done using multiprocessing. Default is False.
- Example
>>> x = BlobLocationComputer(data_path=r"C:/troubleshooting/RAT_NOR/project_folder/videos/2022-06-20_NOB_DOT_4_downsampled_bg_subtracted.mp4", multiprocessing=True, gpu=True, batch_size=2000, save_dir=r"C:/blob_positions") >>> x.run()
Egocentric data / video alignment
- class simba.data_processors.egocentric_aligner.EgocentricalAligner(data_dir: Union[str, PathLike], save_dir: Union[str, PathLike], anchor_1: str = 'tail_base', anchor_2: str = 'nose', direction: int = 0, core_cnt: int = -1, fill_clr: Tuple[int, int, int] = (250, 250, 255), verbose: bool = True, gpu: bool = False, videos_dir: Optional[Union[str, PathLike]] = None, anchor_location: Optional[Union[Tuple[int, int], str]] = (250, 250))[source]
Aligns and rotates movement data and associated video frames based on specified anchor points to produce an egocentric view of the subject. The class aligns frames around a selected anchor point, optionally rotating the subject to a consistent direction and saving the output video.
See also
To produce rotation vectors, uses
egocentrically_align_pose_numba()oregocentrically_align_pose(). To rotate video only, seeEgocentricVideoRotator()- Parameters
config_path (Union[str, os.PathLike]) – Path to the configuration file.
save_dir (Union[str, os.PathLike]) – Directory where the processed output will be saved.
data_dir (Optional[Union[str, os.PathLike]]) – Directory containing CSV files with movement data.
anchor_1 (Optional[str]) – Primary anchor point (e.g., ‘tail_base’) around which the alignment centers.
anchor_2 (Optional[str]) – Secondary anchor point (e.g., ‘nose’) defining the alignment direction.
direction (int) – Target angle, in degrees, for alignment; e.g., 0 aligns east
anchor_location (Optional[Tuple[int, int]]) – Pixel location in the output where anchor_1 should appear; default is (250, 250).
fill_clr (Tuple[int, int, int]) – If rotating the videos, the color of the additional pixels.
rotate_video (Optional[bool]) – Whether to rotate the video to align with the specified direction.
core_cnt (Optional[int]) – Number of CPU cores to use for video rotation; -1 uses all available cores.
- Example
>>> aligner = EgocentricalAligner(rotate_video=True, anchor_1='tail_base', anchor_2='nose', data_dir=r"/data_dir", videos_dir=r'/videos_dir', save_dir=r"/save_dir", video_info=r"C:/troubleshooting/mitra/project_folder/logs/video_info.csv", direction=0, anchor_location=(250, 250), fill_clr=(0, 0, 0)) >>> aligner.run()
Heuristic circling detector
- class simba.data_processors.circling_detector.CirclingDetector(config_path: Union[str, PathLike], nose_name: Optional[str] = 'nose', data_dir: Optional[Union[str, PathLike]] = None, left_ear_name: Optional[str] = 'left_ear', right_ear_name: Optional[str] = 'right_ear', tail_base_name: Optional[str] = 'tail_base', center_name: Optional[str] = 'center', time_threshold: Optional[int] = 10, circular_range_threshold: Optional[int] = 340, shortest_bout: int = 100, movement_threshold: Optional[int] = 60, save_dir: Optional[Union[str, PathLike]] = None)[source]
Detect circling using heuristic rules.
Important
Circling is detected as present when the circular range of the animal is above the specied circular range threshold within the specified preceding time threshold AND the movement of the animal (defined as the sum of the center movement) is above the specified movement threshold within the specified preceding time threshold.
Circling is detected as absent when not present.
Note
We pass the names of the left ear, right ear, and nose, as the method will use these body-parts to compute the direction of the animal in degrees.
- Parameters
data_dir (Union[str, os.PathLike]) – Path to directory containing pose-estimated body-part data in CSV format.
config_path (Union[str, os.PathLike]) – Path to SimBA project config file.
nose_name (Optional[str]) – The name of the pose-estimated nose body-part. Defaults to ‘nose’.
left_ear_name (Optional[str]) – The name of the pose-estimated left ear body-part. Defaults to ‘left_ear’.
right_ear_name (Optional[str]) – The name of the pose-estimated right ear body-part. Defaults to ‘right_ear’.
tail_base_name (Optional[str]) – The name of the pose-estimated tail base body-part. Defaults to ‘tail_base’.
center_name (Optional[str]) – The name of the pose-estimated center body-part. Defaults to ‘center’.
time_threshold (Optional[int]) – The time window in preceding seconds in which to evaluate the animals circular range. Default: 10.
circular_range_threshold (Optional[int]) – A value in degrees, between 0-360.
movement_threshold (Optional[int]) – A movement threshold in millimeters. Defaults to 60.
save_dir (Optional[Union[str, os.PathLike]]) – Directory where to store the results. If None, then results are stored in the
logsdirectory of the SimBA project.
References
- 1
Sabnis et al., Visual detection of seizures in mice using supervised machine learning, biorxiv, doi: https://doi.org/10.1101/2024.05.29.596520.
- 2
Lazaro et al., Brainwide Genetic Capture for Conscious State Transitions, biorxiv, doi: https://doi.org/10.1101/2025.03.28.646066
- Example
>>> CirclingDetector(data_dir=r'D: roubleshooting\mitra\project_folder\csv\outlier_corrected_movement_location', config_path=r"D: roubleshooting\mitra\project_folder\project_config.ini")
Heuristic freezing detector
- class simba.data_processors.freezing_detector.FreezingDetector(config_path: Union[str, PathLike], nose_name: str = 'nose', left_ear_name: str = 'left_ear', right_ear_name: str = 'right_ear', tail_base_name: str = 'tail_base', data_dir: Optional[Union[str, PathLike]] = None, time_window: int = 4, movement_threshold: int = 5, shortest_bout: int = 100, save_dir: Optional[Union[str, PathLike]] = None)[source]
Detect freezing behavior using heuristic rules based on movement velocity thresholds. Analyzes pose-estimation data to detect freezing episodes by computing the mean velocity of key body parts (nape, nose, and tail-base) and identifying periods where movement falls below a specified threshold for a minimum duration.
Important
Freezing is detected as present when the velocity (computed from the mean movement of the nape, nose, and tail-base body-parts) falls below the movement threshold for the duration (and longer) of the specified time-window. Freezing is detected as absent when not present.
Note
The method uses the left and right ear body-parts to compute the nape location of the animal as the midpoint between the ears. The nape, nose, and tail-base movements are averaged to compute overall animal movement velocity.
- Parameters
data_dir (Union[str, os.PathLike]) – Path to directory containing pose-estimated body-part data in CSV format. Each CSV file should contain pose estimation data for one video.
config_path (Union[str, os.PathLike]) – Path to SimBA project config file (.ini format) containing project settings and video information.
nose_name (Optional[str]) – The name of the pose-estimated nose body-part column (without _x/_y suffix). Defaults to ‘nose’.
left_ear_name (Optional[str]) – The name of the pose-estimated left ear body-part column (without _x/_y suffix). Defaults to ‘Left_ear’.
right_ear_name (Optional[str]) – The name of the pose-estimated right ear body-part column (without _x/_y suffix). Defaults to ‘right_ear’.
tail_base_name (Optional[str]) – The name of the pose-estimated tail base body-part column (without _x/_y suffix). Defaults to ‘tail_base’.
time_window (Optional[int]) – The minimum time window in seconds that movement must be below the threshold to be considered freezing. Only freezing bouts lasting at least this duration are retained. Defaults to 3.
movement_threshold (Optional[int]) – Movement threshold in millimeters per second. Frames with mean velocity below this threshold are considered potential freezing. Defaults to 5.
shortest_bout (Optional[int]) – Minimum duration in milliseconds for a freezing bout to be considered valid. Shorter bouts are filtered out. Defaults to 100.
save_dir (Optional[Union[str, os.PathLike]]) – Directory where to store the results. If None, then results are stored in a timestamped subdirectory within the
logsdirectory of the SimBA project.
- Returns
None. Results are saved to CSV files in the specified save directory: - Individual video results: One CSV file per video with freezing annotations added as a ‘FREEZING’ column (1 = freezing, 0 = not freezing) - Aggregate results: aggregate_freezing_results.csv containing summary statistics for all videos
- Example
>>> x = FreezingDetector(data_dir=r'D:\troubleshooting\mitra\project_folder\csv\outlier_corrected_movement_location', config_path=r"D:\troubleshooting\mitra\project_folder\project_config.ini", time_window=3, movement_threshold=5, shortest_bout=100 >>> x.run()
References
- 1
Sabnis et al., Visual detection of seizures in mice using supervised machine learning, biorxiv, doi: https://doi.org/10.1101/2024.05.29.596520.
- 2
Lopez et al., Region-specific Nucleus Accumbens Dopamine Signals Encode Distinct Aspects of Avoidance Learning, biorxiv, doi: https://doi.org/10.1101/2024.08.28.610149
- 3
Lopez, Gabriela C., Louis D. Van Camp, Ryan F. Kovaleski, et al. “Region-Specific Nucleus Accumbens Dopamine Signals Encode Distinct Aspects of Avoidance Learning.” Cell Biology, Volume 35, Issue 10p2433-2443.e5May 19, 2025. DOI: 10.1016/j.cub.2025.04.006
- 4
Lazaro et al., Brainwide Genetic Capture for Conscious State Transitions, biorxiv, doi: https://doi.org/10.1101/2025.03.28.646066
- 5
Sabnis et al., Visual detection of seizures in mice using supervised machine learning, 2025, Cell Reports Methods 5, 101242 December 15, 2025.
Data GPU methods
- simba.data_processors.cuda.data.egocentrically_align_pose_cuda(data: ndarray, anchor_1_idx: int, anchor_2_idx: int, anchor_location: ndarray, direction: int, batch_size: int = 1000000) Tuple[ndarray, ndarray, ndarray][source]
Aligns a set of 2D points egocentrically based on two anchor points and a target direction using GPU acceleration.
Rotates and translates a 3D array of 2D points (e.g., time-series of frame-wise data) such that one anchor point is aligned to a specified location, and the direction between the two anchors is aligned to a target angle.
See also
For numpy function, see
simba.utils.data.egocentrically_align_pose(). For numba alternative, seesimba.utils.data.egocentrically_align_pose_numba(). To align both pose and video, seesimba.data_processors.egocentric_aligner.EgocentricalAligner(). To egocentrically rotate video, seesimba.video_processors.egocentric_video_rotator.EgocentricVideoRotator()EXPECTED RUNTIMES
FRAMES (MILLIONS)
CUDA TIME (S)
CUDA TIME (STEV)
0.25
0.1882001
0.15372434
0.5
0.1221498
0.00574819
1
0.24
0.0307
2
0.38
0.1092
4
0.505
0.01590969
8
1.037
0.0346
16
3.42
1.53194867
7 BODY-PARTS PER FRAME
3 ITERATIONS
batch size 16M
DIDN’T TEST HIGHER N AS I DON’T HAVE THE NON-GPU RAM
- Parameters
data (np.ndarray) – A 3D array of shape (num_frames, num_points, 2) containing 2D points for each frame. Each frame is represented as a 2D array of shape (num_points, 2), where each row corresponds to a point’s (x, y) coordinates.
anchor_1_idx (int) – The index of the first anchor point in data used as the center of alignment. This body-part will be placed in the center of the image.
anchor_2_idx (int) – The index of the second anchor point in data used to calculate the direction vector. This bosy-part will be located direction degrees from the anchor_1 body-part.
direction (int) – The target direction in degrees to which the vector between the two anchors will be aligned.
anchor_location (np.ndarray) – A 1D array of shape (2,) specifying the target (x, y) location for anchor_1_idx after alignment.
batch_size (int) – Size of data that is processed on each iteration on GPU. default 1m. Increase if GPU allows.
- Returns
A tuple containing the rotated data, and variables required for also rotating the video using the same rules: - aligned_data: A 3D array of shape (num_frames, num_points, 2) with the aligned 2D points. - centers: A 2D array of shape (num_frames, 2) containing the original locations of anchor_1_idx in each frame before alignment. - rotation_vectors: A 3D array of shape (num_frames, 2, 2) containing the rotation matrices applied to each frame.
- Return type
Tuple[np.ndarray, np.ndarray, np.ndarray]
- Example
>>> DATA_PATH = r"/mnt/c/Users/sroni/OneDrive/Desktop/rotate_ex/data/501_MA142_Gi_Saline_0513.csv" >>> VIDEO_PATH = r"/mnt/c/Users/sroni/OneDrive/Desktop/rotate_ex/videos/501_MA142_Gi_Saline_0513.mp4" >>> SAVE_PATH = r"/mnt/c/Users/sroni/OneDrive/Desktop/rotate_ex/videos/501_MA142_Gi_Saline_0513_rotated.mp4" >>> ANCHOR_LOC = np.array([300, 300]) >>> >>> df = read_df(file_path=DATA_PATH, file_type='csv') >>> bp_cols = [x for x in df.columns if not x.endswith('_p')] >>> data = df[bp_cols].values.reshape(len(df), int(len(bp_cols)/2), 2).astype(np.int64) >>> data, centers, rotation_matrices = egocentrically_align_pose_cuda(data=data, anchor_1_idx=6, anchor_2_idx=2, anchor_location=ANCHOR_LOC, direction=180,batch_size=36000000)
Light-dark box analysis
- class simba.data_processors.light_dark_box_analyzer.LightDarkBoxAnalyzer(data_dir: Union[str, PathLike], body_part: str, fps: Union[int, float], threshold: float = 0.01, minimum_episode_duration: float = 1e-15, save_path: Optional[Union[str, PathLike]] = None)[source]
Bases:
objectPerform light–dark box analysis using DeepLabCut pose estimation data.
Note
If the specified body-part is detected below the specified threshold, then the animal is the dark-box. Otherwise its in the light-box.
See also
For light/dark box data plotting, see
simba.plotting.light_dark_box_plotter.LightDarkBoxPlotter().This class analyzes animal transitions between light and dark compartments in a light–dark box behavioral test based on the availability of pose-estimation data. It assumes that if pose-estimation for a specified body part is available, the animal is in the light box; otherwise, the animal is in the dark box. It detects bouts (continuous time segments) for each condition and saves the bout-level data to a CSV file.
- param Union[str, os.PathLike] data_dir
Directory containing DeepLabCut CSV files with pose-estimation data.
- param Union[str, os.PathLike] save_path
Full path to save the resulting CSV file with light/dark bouts.
- param str body_part
The name of the body part used to infer the animal’s presence.
- param Union[int, float] fps
Frames per second of the video recordings.
- param float threshold
Value between 0 and 1. If below this value, animal is in dark box. If above, animal is in light box.
- example
>>> python light_dark_box_analyzer.py --data_dir 'D:\light_dark_box\project_folder\csv\input_csv' --save_path "D:\light_dark_box\project_folder\csv
- esultslight_dark_data.csv” –body_part nose –fps 29 –threshold 0.01
>>> analyzer = LightDarkBoxAnalyzer(data_dir='C: roubleshooting wo_black_animals_14bp\dlc_test', save_path="C: roubleshooting wo_black_animals_14bp\light_dark_ex\light_dark_data.csv", body_part='Nose_1', fps=30.2) >>> analyzer = LightDarkBoxAnalyzer(data_dir='D:\light_dark_box\project_folder\csv\input_csv est', save_path="C: roubleshooting wo_black_animals_14bp\light_dark_ex\light_dark_data.csv", body_part='nose', fps=29) >>> analyzer.run() >>> analyzer.save()
- references
- 1
For discussion about the development, see - GitHub issue 446.